Mass photometry characterizes nucleic acids, proteins and their interactions
Interactions between nucleic acids and proteins are critical for many cellular processes. However, protein-nucleic acid interactions often result in highly heterogenous complexes, both in the type of binding partners and in the stoichiometry of the protein-DNA interactions. This heterogeneity presents a challenge to conventional bioanalytical methods.
Mass photometry is a powerful technology that analyzes the mass distribution of the molecules present in a sample at the single-molecule level, without labels or immobilization. Thanks to these strengths, mass photometry provides a detailed overview of all the molecules present in a sample as well as the complexes they form, even when they are present at low abundance.
Measuring protein-DNA affinity with mass photometry vs. SPR
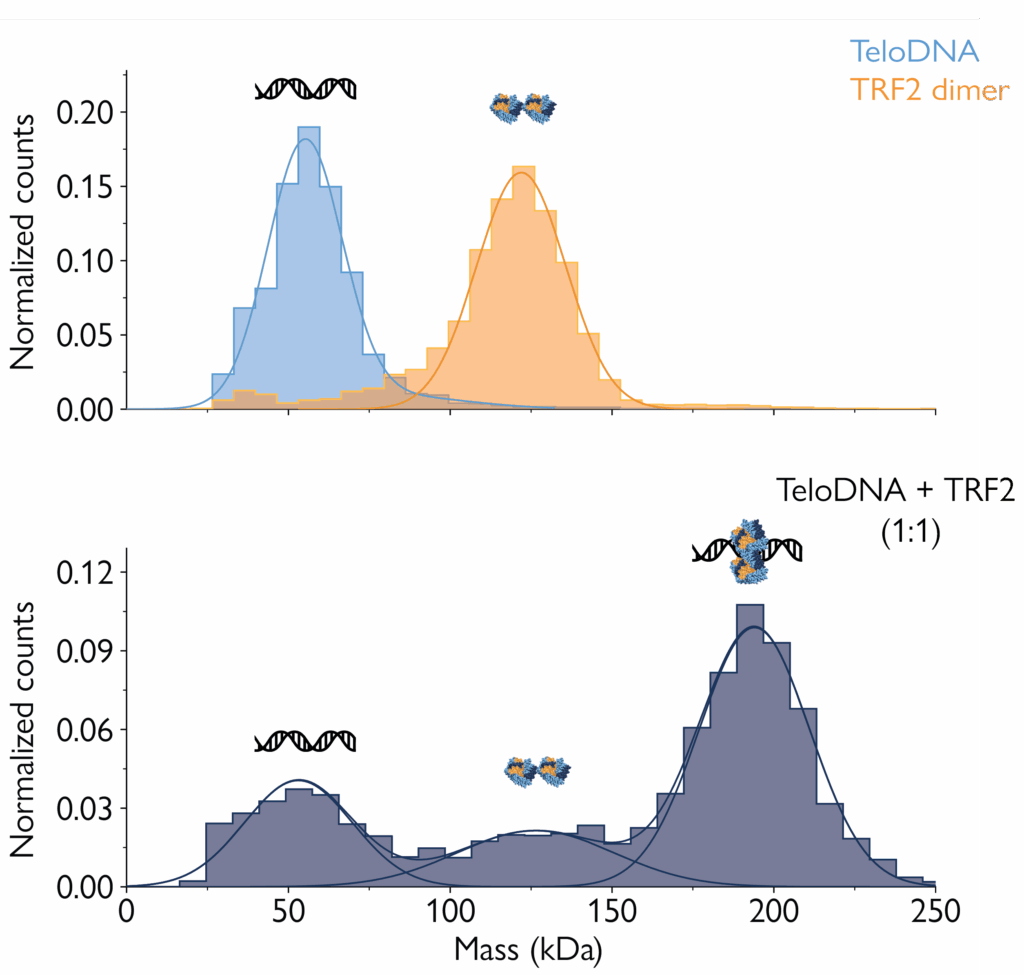
Mass photometry gives information about the proportions of biomolecular species in a sample – making it possible to quantify their binding affinities. As this application note demonstrates, dissociation constants (KD) for nucleic acid-protein complexes calculated from mass photometry measurements agree with results from surface plasmon resonance (SPR) – the analytical method considered the gold standard for assessing protein-DNA binding affinities.
Compared to SPR, mass photometry has the advantages of being faster, not requiring protein immobilization and being simpler to use. Mass photometry also provides valuable information on the stoichiometry of the complexes and relative abundance of the species present in the sample.
To find out how to quickly quantify protein and DNA binding affinities
Characterizing protein-DNA interactions with mass photometry
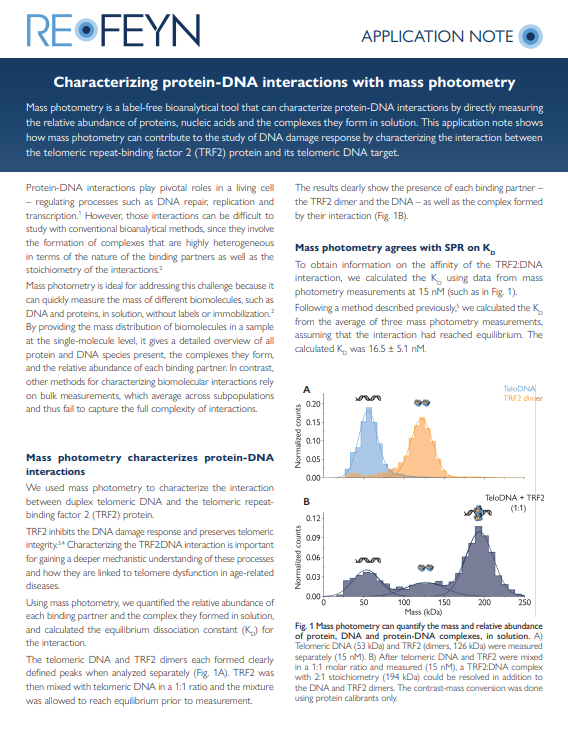
In this application note, you will learn how mass photometry can be used to successfully measure the mass of nucleic acid and protein molecules, and to quantify their abundance and the abundance of complexes they form. You will see data proving that mass photometry can detect DNA and proteins in complex as well as separately, providing the necessary data to quantify their dissociation constant – with results consistent with SPR.
Additional resources on using mass photometry to characterize protein samples
WEBINAR: Elucidating the composition of CRISPR-Cas12f1 complexes using mass photometry
This webinar explores how the CRISPR-Cas12f1 effector complex recognizes and cleaves DNA. Mass photometry was used to measure the stoichiometry of two miniature Cas12f1 ribonucleoprotein complexes, AsCas12f1 and SpCas12f1 in vitro. The results indicate that the Cas protein forms a binary complex with guide RNA and remains bound when interacting with target DNA, forming a ternary complex. Overall, the results reveal the composition of a compact CRISPR-Cas system and its impact on target recognition and DNA cleavage mechanisms.


BLOG: Studying protein-DNA interactions with mass photometry
In this blog post, we present a recent study that used mass photometry to investigate protein-DNA interactions. We then speak with the paper’s first author, Ananya Acharya, who explains how she and her colleagues used mass photometry to answer unique scientific questions about DNA repair mechanisms and compares it to other techniques.
To learn more about how mass photometry can improve your bioanalytical processes
More Application Notes
Browse through our catalogue of application notes highlighting some recent case studies featuring mass photometry.
